Newswise — Karen Lewis, assistant professor in the Department of Chemistry and Biochemistry at Texas State University, has received a $460,000 competitive grant renewal from the National Institutes of Health to study the RNA-to-protein translation process that is controlled by La-Related Proteins (LaRPs).
The three-year grant was awarded by the NIH's National Institute of General Medical Studies and will fund Lewis' project, "Identification of the molecular mechanisms of RNA binding by the posttranscriptional regulator LaRP6."
To express information encoded in genes, living things transcribe the DNA code into messenger RNA molecules, which then instruct the synthesis of proteins. The innovative use of fish protein enables Lewis’ lab to probe the biochemistry, structure, and function of LaRP6, an RNA-binding protein which is shared widely by plants and animals, including human. It is critical to understand the structural and biochemical aspects of these proteins in order to fully understand their biological role. As closer evolutionary relatives to mammals, fish retain considerable similarity to humans that allow Lewis to readily apply discoveries to human systems, while also being sufficiently different to allow identification of functionally important parts of the protein. As fundamental science, discovering the various roles of this poorly understood protein could potentially shed new light on disease development and progression in humans.
“All of the molecular-level work that had been done to date on LARP-6 had been done with human cells or with plant cells. There was a big gap. Plants are very different from humans, and there are many organisms in between,” Lewis said. “To figure out what LARP-6 is doing, we looked at several different species and were able to find fish proteins that are very stable and great to work with. They are providing us some really interesting insight into what LARP-6 is doing.
“This is what we call comparative phylogenetics, where we’re taking the LAPR6 protein from humans and the LARP6 protein from fish, then comparing and contrasting the two to see what’s changed between the two in the protein sequence and the protein structure, what it binds to,” she said. “We’re using those differences and similarities to suss out what this protein does, how it regulates that flow of information from RNA to protein.”
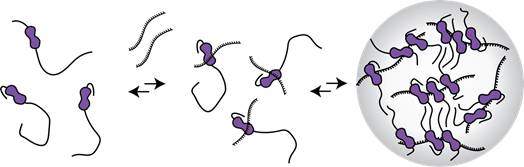
Diagram of how the LaRP6 protein might bind to RNA and then create a complex to regulate translation.
Lewis’ initial grant enabled her team to map three components of LARP6. The first component, N-Terminal Region (NTR), is well-conserved across vertebrates but its functions are largely unknown. The second component, the RNA-binding domain, is the best understood region, as it directly interacts with RNA that leads to the formation of proteins. The third component, however, is poorly understood—at the molecular level, its structure and interactions remain much of a mystery.
“How does that N-terminal region affect the RNA-binding activities of that core domain? We have evidence that it does,” Lewis said. “As far as the biologic and physiologic function, we need to know what it’s doing and why it’s doing it in the context of the whole protein, not just in the isolated binding domain.
“We only know one native target of LaRP6 right now, and that’s collagen. When you disrupt LaRP6 binding to the collagen RNA—the message that is going to make collagen—we know that reduces the amount of collagen that is made. We know that is regulated by LaRP6,” she said. “We want to learn what other proteins LaRP6 is doing that for. LaRP6 is not so specialized that it’s controlling just one protein synthesis. We know that LARP-6 is expressed in cells that don’t express collagen, so why express something if the target’s not there? LaRP6 is expressed in plants and there’s no collagen in plants.
“This is where our comparative phylogenetics comes in. We’re exploring some of the protein structure and function in our lab, and we have a collaboration to figure out what else LaRP6 is regulating in the cell,” Lewis said. “Our expectation is that we’ll be able to link that back to understanding development and disease progression in humans. When there are errors in translation, that’s where you could get disease. Understanding how the cell can regulate protein synthesis at the level of the messenger RNA is important. It’s foundational and fundamental. We’re looking at how the LaRPs do that.”